IMPORTANCE OF PLASMA MEMBRANE IN THE CELLULAR ORIGIN AND EVOLUTION
Atsushi Hase1) and Hakobu Nakamura2)
1. Department of Microbiology, Osaka City Institute of Public Health and Environmental Sciences,
8-34 Tojo, Tennoji, Osaka 543-0026, Japan
Fax:+81-6-6772-0676, Phone:+81-6-6771-3148
E-mail:a.hase@iphes.city.osaka.jp
2. Emeritus Professor of Konan University,
3-2-5-1326 Koyocho Naka, Higashinada, Kobe 658-0032, Japan
(Received 10 April, 2003 Accepted 18 November 2003)
(Abstract)
Life was generated about four billion years ago in the primitive sea as a result of the chemical evolution. The proto-cell was a prokaryotic monad that was formed from a certain aggregate of primitive proteins, nucleic acids, and other macromolecules and wrapped with (phospho)lipid membrane. That membrane served a very important role in the proto-cell born in the primitive sea. This communication addresses the evolutionary process of eukaryotic cell genesis. Eukaryotic cells developed membrane systems remarkably compared to prokaryotic cells. The nucleus, mitochondria, chloroplasts (in plant), endoplasmic reticulum, Golgi apparatus, and other organelles are constructed of lipid membranes inside of which specific enzymes and proteins are positioned. We infer that the eukaryotic organelles were not engendered in an accidental event of symbiotic phenomena, but by evolutionary inevitability of the surviving world. (Key words) cellular evolution, plasma membrane, eukaryotic cell, prokaryotic cell1. Introduction
It has been estimated that life was generated about four billion years ago in primitive sea as a result of chemical evolution. At that time, the proto-organism was a prokaryotic monad: a proto-cell was formed from a certain aggregate of primitive proteins, nucleic acids, and other macromolecules and wrapped with (phospho)lipid membrane. Some investigators have proposed that macromolecules involved in the formation of the life were adsorbed on the surface of clay particles in the primitive soup [1]. However, it seems unlikely that such a simple aggregation of macromolecules would engender formation of a vital cell. A single membrane consisting of a phospholipid bilayer wraps extant cells, without exception. The membrane also played a very important role in the proto-cells birth in the primitive sea because: (a) its cell membrane sets the limit of the vital content to the natural environments; (b), in fact, the phospholipid bilayer does not pass ions. Therefore, for example, it makes a proton (H+) dam in one side of the membrane, which accumulates the potential energy required for various living activities. The lipid membrane can allow selective permeability, by which certain substances involved are more permeable than others. When proteins and ionophores were integrated into the lipid membrane, they produced channels and functioned as specific carriers of organic substances and metallic ions. Active transport systems in the extant plasma membrane can transport particular substances under consumption of ATP even contrary to concentration on one side. Direction of that transport is specific for the membrane protein, permease.The plasma membrane and its derivatives of the extant prokaryotes have been demonstrated to contain enzyme (and proteins) responsible for macromolecular synthesis, photosynthesis, respiration, permeability, and other functions. It may be not too much to say that the prokaryotic plasma membrane contains the most important machineries for life. The lipid membrane plays a role as a kind of the shelf for macromolecules arrangement. On the basis of the cell fossils with large size of eukaryotic level, we inferred that eukaryotic cells originated about 1.5 billion years ago. In addition to the plasma membrane, eukaryotic cells developed membrane systems remarkably in comparison to prokaryotic cells. The nucleus, mitochondria, chloroplast (in plant), endoplasmic reticulum, Golgi apparatus and other organelles are constructed of the lipid membranes, inside of which the specific enzymes and proteins are positioned. The former three organelles contain DNA and are usually enveloped by double membranes comprising inner and outer layers [2]. It is notable that the exclusive era of the prokaryote (about 2.5 billion years) elapsed much longer ago than that of the eukaryotes.
The present communication specifically addresses evolutionary process in the genesis of eukaryotic cells. At present, two hypotheses (endosymbiotic theory and membrane evolution theory) may explain this. The former was re-emphasized by Margulis in 1970 and 1981; Nakamura discussed the latter in 1975. We will explain and examine the contents of these theories in detail in the following sections.
2. Membrane evolution
We can assume that membranous system of the proto-cells comprised only plasma membranes as they do in the present day Mycoplasma. The Mycoplasma cell is the smallest microorganism in size (125-250 nm in diameter) in the biological world and its DNA content is much lower on the average (4.4-12 x 108 in molecular weight) than that of bacterium (8-39 x 108). The genome size (580-2580 kbp) (usually haploid in prokaryotes) is presumed to contain 500-2550 genes, which corresponds the lowest information in prokaryotic and eukaryotic worlds. However, they can grow and multiply in completely artificial media (not in vital cells). On the other hand, Loomis [3] has estimated the genes required to construct the proto-cell as numbering at least 150. Therefore, even the most primitive Mycoplasma cells appear to have evolved considerably in the genome. It is important in the cellular evolution that the plasma membrane of Mycoplasma shows a typical unit membrane structure, but its molecular species of phospholipids is quite simple [4].Archaebacterial cells are phylogenetically unique compared to prokaryota and eukaryota [5]; moreover, their plasma membrane is constructed uniquely by a monolayer of particular lipids. In Gram-positive bacterium Bacillus subtilis, we can observe a membrane complex, called a mesosome, which is derived from the plasma membrane through elongation and folding. The mesosomal membrane shows an oxidation-reduction of redox indicators. Therefore, some investigators refer to this organelle that contains so-called electron transfer and phosphorylation systems as mitochondrion. Therefore they believe it to be a proto-mitochondrion [2,6,7]. On the other hand, mesosomes have been observed to connect with the nucleoid of bacterial cell. Therefore, it is hypothesized that the organelle contains machinery for DNA replication (replicon theory, [8]). Generally, Escherichia coli does not contain an intracellular membrane system except for the plasma membrane. However, we have found that when the E. coli cells were treated with acriflavine (a kind of basic dye and an inhibitor of DNA synthesis through plasma membrane), this drug can effectively eliminate plasmid from the cells. An alkaline broth (pH 8.1) induces formation of a folded membrane system at the end(s) of the rod cell. We isolated an acriflavine-sensitive (acrA) mutant from the wild type (acrA +) strain of E. coli. The acrA gene is located at min 10.6 on the chromosome, just near gene dnaZ for DNA synthesis [9]. Recent work showed that acrA protein is located in a bacterial membrane and plays an important role in the drug efflux system in Gram-negative bacteria [10,11]. Plasmid becomes quite unstable in the mutant cell and is eliminated in the presence of membrane attachable agents in addition to acriflavine and by heating at 42℃. On the other hand, the plasmid is stable in the wild type cell under these conditions [4,12]. Further, induction of the membrane complex by acriflavine occurs more easily and is more complex in the acrA mutant cell than in the acrA + cell. Biochemical analysis showed that acriflavine permits an increased synthesis of phospholipids, particularly phosphatidyl ethanolamine and cardiolipin. These results indicate that intracellular membrane formation is induced according to the physiological conditions [7,9,13]. In fact, as shown in the following, occurrence of such membranous organelles has been known to be conditional in prokaryotic and eukaryotic worlds.
The cell of purple non-sulfur bacterium R. rubrum contains a particular membranous system for photosynthesis: a chromatophore. It is deep-purple colored when photosynthesis is active, whereas the system disappears and the cell is non-colored when photosynthesis is not carried out. Our laboratory found that when the wild type strain of R. rubrum was treated with drug acriflavine in an alkaline broth medium for 3 to 5 h at 30℃, mutants that lose purple color and are therefore incapable of photosynthesis are induced at a frequency of 10 -1 to 10 -5 orders, depending on conditions applied [14]. Notably, the starting wild-type strain of R. rubrum used is marked genetically with a character of streptomycin resistance to avoid contamination. Therefore, of course, the obtained white mutant is also streptomycin-resistant. Kuhl et al. [15] isolated the same type mutant of R. rubrum by treatment with a mutagen, ethylmethane sulfonate. We demonstrated by molecular analysis that the wild type cell contains rather large circular plasmid of ca. 50 kbp (Nakamura and Yokomura, unpublished data). Acetone and methanol (7:2) extraction of pigment from the wild type cell showed clear absorbance for carotenoid at 495 and 525 nm and for bacteriochlorophyll at 775 nm; in contrast, that of the white mutant cell showed no absorbance for either pigments. Furthermore, electron micrographs of the wild type cell, which was cultured under somewhat anaerobic condition, showed that chromatophores are packed into the cell, whereas there are few or no chromatophore in the white mutant cell. Therefore, the plasmid apparently contains genes that are responsible for syntheses of photosynthetic pigments and for formation of chromatophores in the cell. On the other hand, we demonstrated that when the white mutant cell was transformed with the plasmid DNA, the wild type pigments and membranous organelles had the capability of being regenerated. It is interesting that the genes responsible for photosynthesis and morphogenesis in the photosynthetic bacteria are located on plasmid DNA. Recent studies have shown that bacterial DNA, both of the main chromosome and plasmid, bind to its plasma membrane and membranous organelle, such as mesosome [16,17,21]. The replicon theory of Jacob et al. had presumed that replication and division to both poles of DNA are controlled by a machinery that is integrated in the membrane system. Further, the chromatophore fraction was mixed directly and digested with restriction enzyme EcoR1. After washing well, it was hybridized with specific segments of the plasmid DNA as probe. Results showed that the chromatophore-binding DNA was a specific region of the plasmid [18,19]. Similar examples have shown that some specific genes may be involved in the DNA-membrane connection (e.g., in B. subtilis [20]) and each of chromosomes in the eukaryotic cell may be attached to a specific region of its nuclear membrane [21]. The DNA and membrane combination is well known for mitochondrion and chloroplasts. We are accumulating evidence indicating that the acrA mutation of E. coli cell engenders the defect of a specific protein (MW 60,000) in the plasma membrane and instability of plasmids, such as sex (F) and drug resistance (R) factors [22,23]. Furthermore, the acrA mutation decreases frequency of the gene recombination to a few percent of the normal cross [24]. It is reasonable to infer from these results that the acrA+ gene product in the membrane is responsible for stability and gene recombination of the plasmid and the exogenote, respectively. We intend to analyze membranous mechanisms of binding and replication of DNA.
We studied Cyanobacteria that evolved from a line of the photosynthetic bacteria. Thereby, we acquired a new type of the energy metabolism that possesses various sizes of plasmids [25]. Generally, eukaryotic cells develop various intracellular membranes; thereby, they develop metabolic compartmentalization. The unicellular acidothermophilic algal class Cyanidiophyceae attracts interest because it is considered to be at the lowest evolutionary level among the eukaryota. Seckbach [26] and Fredrick [27] have hypothesized that these algae belong to Pre-Rhodophyta and phylogenetically link Cyanobacetria and Rhodophyta. These Cyanidian species are in phylogenetically transitional position linking Cyanobacteria and eukaryotic algae as proposed on the basis of cell morphology, manner of reproduction, and some biochemical components such as storage glucans. Some primitive species, for example, Cyanidioschyzon merolae and Cyanidium caldarium, have almost no "typical" membranous system other than the plasma membrane and the DNA organelles of the nucleus, chloroplasts, and mitochondrion. Furthermore, their chloroplasts seem to be surrounded by a single membrane. Fukuda and Nagashima [28] determined DNA contents of Cyanidium cells using a method of 4,6-diamidino 2-phenylindol (DAPI) staining and fluorescence microscopy including a video-intensified microscopy system. They showed that C. caldarium is much lower in DNA amount per cell than Cyanobacteria, or Anabaena variabilis.
Eukaryotes developed during the course of cellular evolution with various non-DNA organelles, such as endoplasmic reticulum, Golgi apparatus, dictyosome, peroxisome, lysosome, and other intracellular structures. These membrane systems comprised a completed network with increased evolutionary level. The number of mitochondria within a cell can change according to physiological conditions, especially on cell-cycle phases. In a certain phase, all mitochondria are interconnected to form a macro-complex. It produces polymorphic branches and resembles the root system of a tree, as revealed by the electron microscopic figures from re-composed serial thin sections of a single cell. On the other hand, when the macro-complexes of mitochondria were stained with DAPI, the DNA fiber contained therein appeared in a manner similar to a continuous light track of a firefly. However, in other phases of the cell cycle, the complex is divided into many units of individual mitochondria, as seen in regular electron micrographs of the eukaryotic cell. These mitochondrial figures can be observed most clearly in yeast cells. It is incorrect to consider that each mitochondrion is a bacterial symbiont despite beliefs of earlier investigators: the organelle can behave dynamically as the other cellular components.
Similar dynamic figures have also been observed in Euglena chloroplasts [29]. Composite electron micrographs of serial thin sections of a cell show a polymorphic macro-complex of chloroplasts surrounding the nucleus in a certain phase of the cycle. Early observers inferred, based on chloroplast ultrastructural morphology, that the chloroplast is a cyanobacterial symbiont that is similar to mitochondria that previously invaded the host cell, but this is also inaccurate conceptualization. In neither free-living aerobic bacteria nor Cyanobacteria in nature does aggregation of the cells occur according the life cycle as in the macro-complex of organelles. Electron transfer systems for respiratory and photosynthetic metabolism are localized in the inner membranes in mitochondria and chloroplasts. In addition, the carriers are arranged along with reaction flows. Internal mitochondrial membranes are cristae, whereas the chloroplast intra-membranes constitute the thylakoids. Both types function in the same manner to produce energy for ATP and NAD(P)H syntheses, as observed in the purple non-sulfur bacteria and Cyanobacteria [25]. Development of these membrane systems is harmonized with the physiological requirement. For example, yeast cells that have been cultured anaerobically completely lose mitochondrial structures: even their outer membranes are broken. However, when the cells were aerated, those structures are constructed gradually according to activities of the respiratory enzymes. Chloroplasts also lose their interior structures in dark conditions and revert into very small proplastids that possess little or no inner membranes. However, if the plant is illuminated, the proplastids become enlarged and the inner membranes are elongated to produce lamellae or grana. Free-living bacteria and Cyanobacteria never have life cycles similar to the organelles. We must point out that the chloroplast thylakoids are arranged singly in the lower plants, but are joined into two, three, or more bands to compose grana structures as occurred with the evolutionary development of the plant world. Therefore, respiratory and photosynthetic organelles can dynamically change their sizes from 1-3 to 5 µm and larger, their inner membranes go from absent or lower degree of membranes to complex systems, depending on evolutionary, physiological, environmental, and differential parameters. We must stress that the organelle's morphologies are never static: they are dynamic, in metabolic activities. This is the same chain as membranous development with endoplasmic reticlum, Golgi apparatus, and others. In addition, Alberts et al. [30] emphasized that both mitochondrion and chloroplast are quite common membranous constructions and functions. There are interesting movements of the membrane that are specific for animal cells: endo- and exo-cytoses [2]. The former is a phenomenon whereby the plasma membrane invaginates to ingest a drop of liquid (called pinocytosis) or solid particle (called phagocytosis) by wrapping it with the plasma membrane and digesting the incorporated nutrient in the cytoplasm. Phagocytosis is a more complex movement than pinocytosis: the vesicle (phagosome) taken in is conjugated with lysosome, which contains various hydrolytic enzymes, to digest the phagosomic content. Exocytosis is the reverse phenomenon of endocytosis: the cytoplasmic vesicles are transferred to the plasma membrane and then eventually conjugate to excrete their content outside of the cell. Such membrane movement is carried out actively and widely in secretory cells to secrete hormones and exoenzymes, and in the excretory cells to excrete fecal and other wastes. An interesting problem in exocytosis arises: when the vesicles conjugate continuously with the plasma membrane, the membrane size, i.e., the cell, must be enlarged. However, the cell size does not increase steadily. This means that the exocytotic membrane is degraded concomitant with the enlargement; thereby a dynamic turnover occurs between the plasma membrane and, perhaps, the Golgi apparatus.
As stated above, the membrane skeleton is a phospholipid bilayer. Therefore, it is very thin and flexible in a single cell and organelle. This suggests that the membrane can expand extensively within such a small space, especially by pleating. Therefore, the membranous structure is quite useful to arrange the integrated proteins, mainly enzymes, and to fix them three-dimensionally in the cytoplasm. Furthermore, the broken membranes are easily aggregated. The membranes are largely responsible for so-called metabolic order in the cell.
3. Symbiotic theory on the origin of eukaryotic cell and its problems
3.1 Symbiotic theoryThe idea of (endo)symbiotic origin of organelles, e.g., mitochondria and chloroplasts (in plant) within the extant eukaryotic cell was presented in the late 1800s. It was derived from the microscopic observation that organelles shapes are analogous to rod-shaped bacteria and unicellular Cyanobacteria, respectively. Margulis [31,32] re-emphasized this idea and also contended that flagella are symbiotic derivatives of the spiral bacterium, Spirochaeta. However, that study did not scrutinize the origin of the nucleus even though it is at the root of the problem. Some symbiotic theory supporters stress that DNA-containing organelles, e.g., mitochondria, chloroplasts, and the nucleus, possess double (inner and outer) membranes. Therefore, they believe that the inner membranes of mitochondria and chloroplasts are derived from corresponding prokaryotic membranes (of endosymbiont), whereas the outer membranes are from the plasma membrane of the host cell. In other words, the endosymbiotic prokaryote was wrapped by the host's own membrane when it invaded; then the cell wall of the invader disappeared. However, according to recent evidence, composition, and therefore function, differs greatly for inner and outer membranes. That is, the outer membrane as the host's plasma membrane contains a large channel-forming protein (called porin), which is a constituent of the cell wall of prokaryotes, e.g., Gram-negative bacteria, and cannot be found in the plasma membrane. Other proteins in this outer membrane include enzymes involved in specific mitochondrial lipid synthesis. On the other hand, the inner membrane is folded into numerous cristae and contains the electron-transfer system. Similar differentiation between inner and outer membranes also occurs in chloroplasts. Whichever the case may be, the outer membranes of these organelles are not the plasma membrane origin of the host cell.
The biological world presents many examples that do not fit this idea: a single in Cyanidium and triple membranes in Euglena and Dinophyceae surrounding chloroplasts; quadruple membranes enclosing chloroplasts in Cryptophyceae, Phaeophyceae, Bacillariophyceae, Chrysophyceae, and Xanthophyceae; a single membrane of the nucleus in Noctiluca. Interestingly, Jensen (1994) has shown, for Oscillatoria and others, that a single thylakoidal membrane surrounds the nucleoid, suggesting eukaryotic differentiation in the cyanobacterial world.
The symbiotic theory explains processes of generation of mitochondria, chloroplast, or flagella simply by mosaic designing of prokaryotes with specific characters corresponding to the organelles. It is generally said that the simpler the theory, the better its acceptance. However, we must bear in mind that the simplicity of theory is independent of the fact; rather, such simplicity engenders greater complexity when explaining details. Symbiotic theory is such a case: it complicates the explanation of the genetic differentiation from the four kinds of symbiotic prokaryotes to a single eukaryotic cell. In fact, it is impossible to explain genetic differentiation by the above theory, at least by the current of genetics. Some advocates of the symbiotic theory describe cellular evolution from prokaryote to eukaryote as a great leap. However, we do not know of such an example of positive (progressive) jumps in the biological world despite our knowledge of examples of negative (extinguished) jumps. Positive evolutions can even be considered to represent the repeated accumulations of minor mutations and selections. Macroevolution in morphology is also a result of accumulation of the microevolution in genetics.
3.2 Rebuttal against symbiotic theory
Some critical comments on the main problems within the symbiotic theory are given here.
3.2.1 Organelle morphology and its internal genomic evolution
The first problem is whether mitochondria and chloroplast have bacteria- and Cyanobacteria-like morphologies, respectively. The figures that pioneers of cytology observed may appear on a single thin section of the cell. If they had observed many figures taken from serial thin sectioning and reconstituted them in a three-dimensional structure, they would have seen that an organelle's actual morphology appears as more complex then that of the corresponding prokaryotes. Moreover, if they had observed the stereo structure through the progressive cell cycle, then the organelle would have been indicated as more dynamic behaviors. Such behavior depends on physiology of the cell where the organelles assemble in one phase of the cycle, but disjoin in the other, as discussed above.
All organelles, whether prokaryote and eukaryote, are products of differentiation along cellular evolution. The increase in the number of certain genes by duplication may be followed by their expression in producing a variety of many enzymes and proteins through accumulation of mutations and selections. Cells exhibit a persistent tendency to increase their metabolic complexity. The organelle plays an important role in compartmentalizing metabolism according to the functional unit. That is, the organelle guarantees specificity and velocity of metabolism. Cellular evolution from prokaryote to eukaryote follows the same line of metabolism compartmentalization through membranous structures because membranous differentiation occurs even within prokaryotic and the eukaryotic worlds, from simple to complex.
Biological morphology is essentially discontinuous in all cells and organisms. However, we know at present that, in spite of such discontinuity in morphology, DNA sequence evolution is largely a continuous process. Apparently, this results from the fact that a morphological change is derived by a minor alteration of the nucleotide sequence. For example, a gorilla differs obviously in external morphology from a human. However, there is little difference in protein sequences between them; it is even said that there might be no difference between them, except for culture. Recently, genetic study of morphologenesis is active, especially through homeotic transformations: genes involved in determining flower structure and in segmenting animal body [33].
3.2.2 Organellar DNA sequence, products and plasmids
Sequencing of whole DNA in mitochondrion and chloroplast performed by Anderson et al. [34], Shinozaki et al. [35], Matsubayashi et al. [36] and subsequent investigators have clarified the roles of genes responsible for morphogenesis and functions of these organelles. It shows first that almost all related content of genes within the nuclear DNA and their products are exported from cytoplasm to the corresponding organelle. Organelle DNA codes for its own tRNAs, rRNAs, small numbers of enzymes and proteins only, and syntheses of some subunits of enzymes including cytochrome oxidase, cytochrome b and ATPase in mitochondria. Coding for subunits of the photosynthetic enzyme ribulose 1,5-bisphosphate carboxylase (Rubisco) in the chloroplast is also shared between the cytoplasm and the organelle itself.
Interestingly, mitochondrial codons differ from each other and from universal codons. Alberts et al. [30] explain this discordance that the mitochondrial DNA encodes some proteins, which erred in their amino acid sequences. Nenertheless, their small synthesized amounts allow cell survival. This explanation means that the universal code of the nuclear DNA is the survivor of a strict selection for cell life. Second, it has been shown that there are homologous sequences among DNA of the mitochondrion, chloroplast, and nucleus; some of them are high in the homology, whereas others are low. Cytochromes b/f (c) in both mitochondrion and chloroplast are examples [34,36]. These facts suggest that some nucleotide sequences duplicated repeatedly before division of DNA among mitochondria, chloroplast, and nucleus. Some believers of the symbiotic theory may assert the occurrence of repeatedly large-scale, but no minor, genetic redistributions among DNA organelles. However, no evidence has been demonstrated and none of the mechanisms have been hypothesized. In fact, no example show such sequence transfer in large scale between DNA of the symbiont and the host and between extant organelles. Although some unfunctional genes may be lost from the symbiont from lack of selective advantage, no evidence shows that the lost gene from the symbiont was picked up and integrated into the host DNA.
An intron is a particular nucleotide sequence that is embedded in some eukaryotic genes and is trimmed off by a splicing enzyme before transcription. Introns have been found not only in nuclear genes but also mitochondrial and chloroplast genes. However, they do not exist in prokaryotic genes (except some archaebacterial species). Some investigators explain this fact as follows: In the early time, prokaryotic genes also contained introns, but lost them during a long evolution. On the other hand, eukaryotic genes maintained them up to the present. Such an explanation contains a serious contradictory concept: all introns would have been eliminated from the entire prokaryotic world simultaneously, a fact that is impossible to reason. A conceivable explanation is that introns were produced during eukaryotic evolution from a prokaryotic line. That is because eukaryotic DNA contains many duplicated sequences and the intron works to partition the primitive genes, the duplication of which engendered the large sequence of the present gene. Therefore, it is quite difficult to consider mitochondrion and chloroplast DNA as a direct-descendant product of prokaryotic corresponding genes despite the positive approach to this by symbiotic advocates.
3.2.3 Gene sequences of bacteria and organelles
Symbiotic proponents assert that some gene sequences of mitochondrion and chloroplast closely match those of corresponding aerobic bacteria and Cyanobacteria, respectively. However, stark facts indicate that gene sequences of non-photosynthetic bacteria are also similar with those of chloroplasts and Cyanobacteria. For example, the sequence of 16S rRNA in the maize chloroplast is highly similar with that of E. coli; also, sequence of 16S rRNA in Rhodophyceae's chloroplast is similar not only to Cyanobacteria, but also with aerobic bacterium, E. coli and B. subtilis. Notwithstanding, there are many dissimilar sequences of genes of the prokaryote and the eukaryotic organelle. The second important fact is that the prokaryotic DNA sequence is also enclosed, in many cases, within the nucleus [3]. A meaningful point in the molecular evolutionary consideration is that DNA sequence homology depends largely on the physiological significance of the sequence. This is true in the whole sequence of the gene and the partial sequence within gene. The more important the gene or its higher level of activity (or its activity center), the more conservative its sequence. The reason is that accumulated mutations in an important gene or a central part of a gene must lead to a disadvantage of the organism in natural selection. This is the same reason that the universal code of prokaryotic and eukaryotic (main) DNAs is very conservative: it underscores the meaning of "universal", as discussed above.
3.2.4 Ribosomal size and protein content
Symbiotic supporters assert that mitochondrial and chloroplast ribosomes have a sedimentation coefficient of prokaryotic type 70S, whereas cytoplasmic ribosomes are non-prokaryotic 80S. However, in fact, there are many various ribosome coefficients; they range from 55S (mammals) to 80S (yeast) in mitochondrial ribosome and from 66S (spinach) to 78S (maize) in chloroplast ribosomes. Cytoplasmic ribosomes range from 77S (Neurospora) to 83S (mammals), depending on the organism. Moreover, the variety of the sedimentation coefficient is remarkable in each of small and large subunits of ribosomes among species of organisms. Some ribosomes are deficient in 5 or 5.8 rRNA. The number of proteins comprising a single ribosome is distributed in the range of 50 to 107 according to the organism.
The cytoplasmic type of ribosome is generally larger than the prokaryotic type, but the former must be derived from the latter. Therefore, evolutionary interest is rather to analyze how the organelle evolves to compose the larger one. After all, ribosomal size cannot be determinative for deducing the synthetic origin of mitochondrion and chloroplast.
3.2.5 Genomic content and intercellular genetic exchange
If the organellar DNA indeed descended from a symbiotic origin, then the eukaryotic cell must first have become a tetraploid genetic mosaic comprising symbionts, aerobic bacterium (mitochondrion), Cyanobacteria (chloroplast), Spirochaeta (flagellum), and Mycoplasma (nucleus) through complex recombination and deletion of their various genes. Symbiotic advocates have not offered any mechanism addressing that problem. Furthermore, such a scenario is very unlikely to have occurred because: (a) genetic cross is not observed beyond species of different kingdoms such as Eubacteria (Bacteriomycota), Cyanobacteria (Cyanophyta), and Mycoplasma (Mycoplasmomycota, presumed as the host for the symbionts) [31]; (b) when a phylogenetically different line of DNA invades a host cell, restriction endonuclease of the latter must digest the invader to preserve its own phyletic line. All biological lines have been maintained by such a mechanism. There is no evidence to show that genetic exchange occurs between the symbiont and the host during their symbiosis.
3.2.6 tRNA species in mitochondria and bacteria
The number of tRNA species involved in mitochondrial synthesis is much lower than that in the bacterial one: mitochondrion usually contains 22-24 species of tRNA, whereas a bacterial cell possesses two to three times more species than the organelle. Furthermore, there are also large differences between mitochondrion and bacteria in codon usage. Therefore, it is difficult to consider that the mitochondrion is a direct symbiont of the bacteria, as in the symbiotic theory.
3.2.7 Status of cyanelles, Cyanobacteria, and chloroplasts
The protistan, Cyanophora paradoxa, is classified on the one hand into Protozoa (animal kingdom) and on the other into Cryptophyta or Glaucophycophyta, within algal divisions. This photosynthetic unicell contains cyanobacterial symbionts in its cytoplasm. The symbiont is distinctively called a "cyanelle" to mean that it is at an intermediate state from Cyanobacteria to chloroplast. The main reason is that the cyanelle's DNA content is 10 % that of a typical Cyanobateria and is comparable to that of a chloroplast in a higher plant. However, the cyanelle has a cyanobacterial cell wall and possesses both types of ribulose 1,5-bisphosphate carboxylase, Rubisco, subunits (eight large and eight small). Rubisco is a gate enzyme of CO2 fixation in the reductive pentose phosphate cycle. Its specific genes on a single DNA molecule, as in Cyanobacteria, determine synthesis. As described in the foregoing section, synthesis of large subunits is determined by chloroplast DNA, whereas that of the small subunits is determined by the nuclear one in higher plants and some algae. Furthermore, it is said that an isolated cyanelle can grow without the host. These facts indicate that the cyanelle is the Cyanobacterium Cyanocyta korschikoffiana that once invaded a host [2]. The cyanobacterial world has adaptively radiated into quite wide phyletic lines and their DNA contents are also known to range from 1.2 x 109 Da in a unicellular species to 1010 Da in a multi-cellular species. Therefore, it is reasonable to infer that the cyanelle C. korschikoffiana is a self-growing unicellular species of Cyanophyta and is not an example in a modified way of chloroplast. It is natural that there are similar DNA sequences in the cyanelle to those in chloroplasts because the latter contains some DNA sequences that are homologous with Cyanobacteria [2,37].
3.2.8 Origin and physiology of chloroplasts and mitochondria
The chloroplast of higher plant cells is known to change largely in morphology and function depending on environmental and physiological conditions. Chloroplasts are one type of general plastid (also encompassing chromoplasts). They are: (a) surrounded by double membranes, (b) contain specific DNA and proteins, and (c) play various roles in photosynthesis and storage of starch and other substances. Additionally, (d) the chloroplast contains various metabolisms that synthesize amino acids, lipids, and others. In animal cells, these substances are synthesized in the cytoplasm. Dark-grown plants and non-illuminated cells within light-grown plants have semi-autoreproducible proplastid - quite small DNA organelles - but contain little or no thylakoid and photosynthetic pigment. When a dark-grown plant is illuminated, the proplastid develops into a matured chloroplast that can actively photosynthesize. Although the symbiotic theory has stress that the chloroplast is a symbiotic derivative of Cyanobacteria, the free-living Cyanobacterium does not possess such the proplastid phase within its life cycle. Currently, there is a hypothesis that a mitochondrion can also develop from promitochondrion according to cell physiology. If this is the case, then aerobic bacteria as the ancestor of mitochondrion has no special phase of promitochondrion in the life cycle.
4. Membrane evolution theory
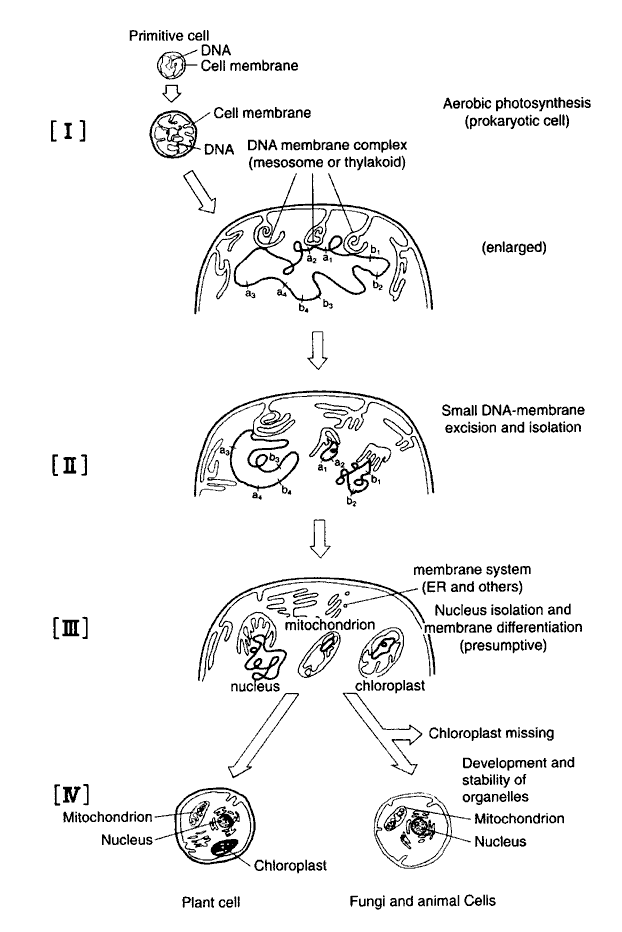
We have inspected and analyzed the content of the symbiotic theory on the origin of the eukaryotic cell. Consequently, we have concluded that the theory does not reasonably explain the eukaryotic cell origin process. Therefore, we proposed a "Membrane Evolution Theory" in 1975 to replace the symbiotic theory of Margulis [32,34]. We now discuss our theory that has been reinforced with new knowledge. It has been established that purple non-sulfur bacteria including R. rubrum contain both metabolisms of the (ancestral) photosynthesis and O2-respiration [6,13]. Similarly, Cyanobacteria, for example C. fritschii, which was derived from photosynthetic bacteria and is believed to link phylogenetically between unicellular and simple filamentous forms [37], provides more evolved photosynthesis which consists of a serial photosystem I and II, evolving O2 gas. Furthermore, the species has a complete O2-respiration metabolism that consists of an EM pathway, a citric acid cycle, and a respiratory chain [38]. In Cyanobacteria, the cell DNA content has been demonstrated to increase duplicatively from unicellular to filamentous form [39]. As indicated above, the evolved species of Cyanobacteria have been endowed with genes coding for eukaryotic photosynthesis and respiration [3,40,41]. Therefore, it is reasonable to infer that the DNA-carrying organelles, such as mitochondria, chloroplasts, and nuclei, were generated up to about 1.5 billion years ago through genetic and membranous differentiation from a line of evolved Cyanobacteria at a similar stage when there occurred differentiation of endoplasmic reticulum, Golgi-apparatus, and other membranous organelles in the pro-eukaryotic cell [42]. However, it is unnecessary to presume that photosynthesis and respiration genes were derived from some separate organisms, which once penetrated into an unknown prokaryotic host. Originally, the cell does not accept foreign DNA and does not allow recombination between DNAs of different species. Species-specific restriction enzymes are primarily so-called arms, which defend the cell against the invader DNA. Therefore, all biological species must have specific enzymes that digest foreign DNA. We must stress here that such a careful mechanism preserves phylogeny in the biological world. In fact, no natural hybridization occurs without a single, or near, taxonomic species. Therefore, genetic recombination occurs only rarely between organisms beyond the species; moreover, it is impossible to combine organisms different Phyla despite the assertion of symbiotic theory that it can occur. (In contrast, symbiosis or parasitism occurs naturally and is popular beyond the taxonomy or even between animal and plant kingdoms).Figure 1 illustrates points of our agreement stated above together with other biological phenomena. A long DNA fiber in the evolved Cyanobacterium was broken into at the three points: two small fragments and one large one. Then, an intracellular membrane, such as thylakoid and mesosome, wrapped the DNA fragments. A small DNA containing some genes relating to citric acid cycle, respiratory chain, t- and r-RNAs, and other components, were organized as the mitochondrion. Another DNA fragment containing some genes relating to reductive pentose phosphate cycle, photosystem, t- and r-RNAs, and other constituents were organized into a chloroplast. Further, the remainder of these DNAs is located in the nucleus. We have inferred that t-and r-RNAs coded for by the mitochondrial and chloroplast DNAs are duplicative offspring that have an ancestral nucleotide sequence common with nuclear t-and r-RNAs, but they diverged phylogenetically concomitant with differentiation. When differentiated cells containing the presumed chloroplasts lost their plastidal genes, they became non-photosynthetic as animal and fungus organisms. Spontaneous or induced, but not lethal, elimination of chloroplasts from green algae has been observed in laboratory environments, e.g. in Chlorella and Euglena.
5. Conclusion
Because the proto-cell was first generated in the primeval soup about four billion years ago, its offspring continued to evolve a cellular metabolism and organelle and multi-cellular morphology throughout prokaryotic and eukaryotic worlds. Such a developmental course to raise the efficiency of life resulted naturally from repeated mutations and selections. Mutations always occur by spontaneous and induced mistakes of base-pairing in DNA. Biological and non-biological conditions in nature also select some organisms that adjust well in their habitat. In other words, cells have always been compelled to adapt their function through metabolic and morphological alterations. Generation of a new organelle and its development to engender enhanced cellular function is an essential process for life; if that process does not occur, then the organism is selected against. Therefore, we conclude that eukaryotic organelles were not born of an accidental event of symbiotic phenomena, but by evolutionary inevitability of the surviving world.Eukaryotic cell genesis has largely advanced by membrane system development. However, the latter has developed with functional complexity along with prokaryote evolution. The membrane system plays a very important role in metabolic compartmentalization. Therefore, we can observe that membranous organelles increase morphological, namely functional, complexity and especially with evolution even in the eukaryotic world. On the other hand, basic metabolisms for life are well preserved throughout prokaryotes and eukayotes, but their efficiency must be raised with morphological evolution. The base sequence of chloroplast DNA coincides completely among liverwort (Bryophyta) in lower plants [43] and tobacco (Corolliforiidae)[35] and rice (Glumifordiidae)[44] in higher plants. Considered together with other data, we infer that DNA of the organelle has specific conservativity throughout the plant kingdom. If symbiotic origin of eukaryotic organelles was usual in the past, then we may be able to observe various types of mitochondria, chloroplasts, and other DNA-carrying eukaryotic organelles in the present. Otherwise, extant eukaryotes must be explained as derivatives of a single ancestor that was selected for among the organelle-heterogenic eukaryotes. Recently, evidence has accumulated to show that the amino acid sequence exist in enzymes that are common to all cells from Eubacteria to higher organisms. The origin of this sequence might have emerged in a chemical evolution that once occurred in the primeval soup [3]. Organisms fundamentally have a strong compulsion to preserve, rather than mix, their phylogeny. The biological mechanism and chemical stability of DNA have accomplished such a defense of genetics, but many problems remain to be illuminated concerning our membrane evolution theory [6,18,33,45,46].
References
1. Cairns-Smith, A. G., Genetic takeover and the mineral origins of life, Cambridge University Press, Cambridge, 1982.2. Nakamura, H., Cellular evolution, Baifukan Publisher, Tokyo, 1987.
3. Loomis, W. F., Four billion years, Sinauer Associated Inc., Sunderland, MA, 1988. 4. Nakamura, H., Membranous evolution in the process of chemical evolution to cellular evolution, Viva Origino 4, 43-61 (1975).
5. Woese, C. R., Dugre, D. H., Dugre, S. A., Kondo, M. and Saxinger, W. C., On the fundamental nature and evolution of the genetic code. Cold Spring Harb. Symp. Quant. Biol. 31, 723-36 (1966).
6. Nakamura, H., Origin of eukaryota from cyanobacterium: membrane evolution theory, In Evolutionary pathways and enigmatic Algae, Seckbach, J. Ed.; pp.3 Kluwer Academic Publishers, Amsterdam, 1994.
7. Nakamura, H., Tsukuba university report of special research project on evolution of matter, vol. 2; pp. 185 (1988).
8. Jacob, F., Brenner, S. and Cuzin, F., On the regulation of DNA replication in bacteria, Cold Spring Harb. Symp. Quant. Bio. 28, 329-348 (1963).
9. Nakamura, H., Gene-controlled resistance to acriflavine and other basic dyes in Escherichia coli, J. Bacteriol. 90, 8-14 (1965).
10. Okusu, H., Ma, D. and Nikaido, H., AcrAB efflux pump plays a major role in the antibiotic resistance phenotype of Escherichia coli multiple-antibiotic-resistance (Mar) mutants, J. Bacteriol. 78, 306-308 (1996).
11. Zgurskaya, H. I. and Nikaido, H., AcrA is a highly asymmetric protein capable of spanning the periplasm, J. Mol. Biol. 285, 409-420 (1999).
12. Nakamura, H., Tojo, T. and Greenberg, J., Interaction of the expression of two membrane genes, acrA and plsA, in Escherichia coli. J. Bacteriol. 122, 74-879 (1975). 13. Nakamura, H., Genetic background of cellular evolution, Mem. Konan. Univ. Sci. Ser. 29, 55-73 (1983).
14. Nakamura, H. and Yoshikawa, Y., Non-purple mutants of purple non-sulfur bacterium Rhodospirillum rubrum, Mem. Konan Univ. Sci. Ser. 33, 1-8 (1986).
15. Kuhl, S. A., Nix, D. W. and Yoch, D. C., Characterization of a Rhodospirillum rubrum plasmid: loss of photosynthetic growth plasmidless strains, J. Bacteriol. 156, 737-742 (1983).
16. Nakamura, H., In Origin of Life, Noda, H., Ed.; pp. 515, Center for Academic Publication, Tokyo, 1978.
17. Kornberg, A., DNA replication, W. H. Freeman and Company, San Francisco, CA, 1980. 18. Nakamura, H., Mem. Konan Univ. Sci. Ser. 37, 87 (1990).
19. Nakamura, H., Metabolic and membranous differentiation leading to non-symbiotic origin of eukaryotic cell, In Endocytobiology V., Sato et al. Ed.; pp. 337, Tubingen University Press, Tubingen, 1993.
20. Yoshikawa, H., DNA synthesis during germination of Bacillus subtilis spores, Proc. Natl. Acad. Sci. USA 53, 1476-1483 (1965).
21. Berns, M. W., Cells (2nd Ed.), CBS Publishing, Los Angels, CA, 1983.
22. Nakamura, H., Plasmid-instability in acrA mutants of Escherichia coli K12, J. Gen Microbiol. 84, 85-93 (1974).
23. Nakamura, H., R factor instability in the acriflavine-sensitive mutant of Escherichia coli K12, Jpn. J. Genet. 51, 393-395 (1976).
24. Nakamura, H., Suganuma, A. and Greenberg, J., Effect of inorganic phosphate on acridine inhibition and plasmid curing in Escherichia coli, J. Gen. Microbiol. 91, 45-52 (1975). 25. Carrn, N. C. and Whitton, B. A., The Biology of Cyanobacteria, University of California Press, Berkeley, CA, 1982.
26. Seckbach, J., Ann. New York Ac. Sci. 503, 424 (1987).
27. Frederick, J. F., Ann. New York Ac. Sci. 503, 438 (1987). 28. Fukuda, I. and Nagashima, H., Ann. New York Ac. Sci. 503, 575 (1987).
29. Ehara. T., Osafune, T. and Hase, E., Interactions between the nucleus and cytoplasmic organelles during the cell cycle of Euglena gracilis in synchronized cultures. IV. An aggregate form of chloroplasts in association with the nucleus appearing prior to chloroplast division, Exp. Cell Res. 190, 104-12 (1990).
30. Alberts, B., Molecular biology of the cell (3rd ed.), Garland Publishing, New York, NY, 1994.
31. Margulis, L., Origin of eukaryotic cell, Yale University Press, New Haven, CT, 1970. 32. Margulis, L., Symbiosis in cell evolution ミLife and its environment of the early earth, W. H. Freeman and Company, San Francisco, CA, 1981.
33. Nakamura, H., Basic biology-Life mechanism on the molecular and cellular aspect, Baifukan Publisher, Tokyo, 2000.
34. Anderson, S., Bankier, A. T., Barrell, B. G., de Bruijin, M. H., Coulson, A. R., Drouin, J., Eperon, I. C., Nierlich, D. P., Roe, B. A., Sanger, F., Schreier, Smith, A. J, Standen, R. and Young, I. G., Sequence and organization of the human mitochondrial genome, Nature 290, 457-465 (1981).
35. Shinozaki, K., Deno, H., Wakasugi, T. and Sugiura, M., Tobacco chloroplast gene coding for subunit I of proton-translocating ATPase: comparison with the wheat subunit I and E. coli subunit b, Curr. Genet. 10, 421-3 (1986).
36. Matsubayashi, T., Wakasugi, T., Shinozaki, K., Yamagichi-Shinozaki, K., Zaita, N., Hidaka, T., Meng, B. Y., Ohto, C., Tanaka, M. and Kato, A., Six chloroplast genes (ndhA-F) homologous to human mitochondrial genes encoding components of the respiratory chain NADH dehydrogenase are actively expressed: determination of the splice sites in ndhA and ndhB pre-mRNAs. Mol. Gen. Genet. 210, 385-393 (1987).
37. Fogg, G. E., In The Blue-Green Algae, Academic Press, London, 1973. 38. Ragan, M. A. and D. J. Chapman, A Biochemical Phylogeny of the Protists, Academic Press, New York, NY, 1978.
39. Herdman, M. and Carr, N. G., The isolation and characterization of mutant strains of the blue-green alga Anacystis nidulans, J. Gen. Microbiol. 70, 213-20 (1972).
40. Peschek, G. A., Strcture and function of respiratory membranes in Cyanobacteria (blue-green algae), Subcell Biochem. 10, 85-191 (1984).
41. Peschek, G. A., In Cyanobacteria, Fay, P. and van Baaken, C. Ed., Elsevier, Amsterdam, 1987.
42. Sere, P. A. and Estabrook, R. W, Microenvironments and metabolic compartmentation, Academic Press, New York, NY, 1978.
43. Ohyama,K., Fukuzawa, H., Kohchi, T., Shirai, Sano, T., Sano, S., Umesono, K., Shiki, Y., Takeuchi, M., Chang, Z., Aota, S., Inokuchi, H., and Ozeki, H., Chloroplast gene organization deduced from complete sequence of liverwort Marchania polymorpha chloroplast DNA, Nature 322, 572-574 (1986).
44. Hiratsuka, J. Shimada, H., Whitter, R., Ishibashi, T., Sakamoto, M., Mori, M., Kondo, C., Honji, Y., Sun, C. R. and Meng, B. Y., The complete sequence of the rice (Oryza sativa) chloroplast genome: intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of the cereals, Mol. Gen. Genet. 217, 185-94 (1989).
45. Nakamura, H. and Hase, A., Cellular differentiation in the process of generation of the eukaryotic cell, Origin of Life and Evolution of the Biosphere 20, 499-514 (1990).
46. Nakamura, H., In Origin and Evolution of the Cell, Hartman, H. and Matsuno, K., Ed., World Scientific, Singapore, 1992.